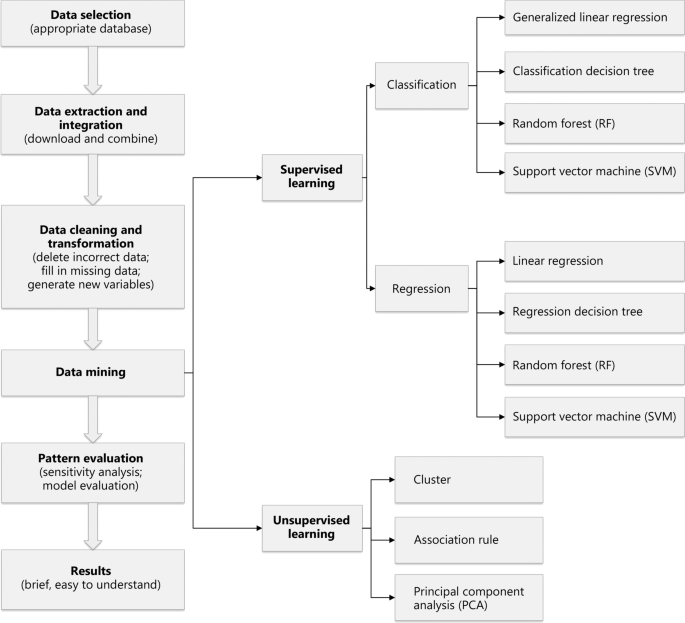
Many high quality studies have emerged from public databases, such as Surveillance, Epidemiology, and End Results (SEER), National Health and Nutrition Examination Survey (NHANES), The Cancer Genome Atlas (TCGA), and Medical Information Mart for Intensive Care (MIMIC); however, these data are often characterized by a high degree of dimensional heterogeneity, timeliness, scarcity, irregularity, and other characteristics, resulting in the value of these data not being fully utilized. Data-mining technology has been a frontier field in medical research, as it demonstrates excellent performance in evaluating patient risks and assisting clinical decision-making in building disease-prediction models. Therefore, data mining has unique advantages in clinical big-data research, especially in large-scale medical public databases. This article introduced the main medical public database and described the steps, tasks, and models of data mining in simple language. Additionally, we described data-mining methods along with their practical applications. The goal of this work was to aid clinical researchers in gaining a clear and intuitive understanding of the application of data-mining technology on clinical big-data in order to promote the production of research results that are beneficial to doctors and patients.
With the rapid development of computer software/hardware and internet technology, the amount of data has increased at an amazing speed. “Big data” as an abstract concept currently affects all walks of life [1], and although its importance has been recognized, its definition varies slightly from field to field. In the field of computer science, big data refers to a dataset that cannot be perceived, acquired, managed, processed, or served within a tolerable time by using traditional IT and software and hardware tools. Generally, big data refers to a dataset that exceeds the scope of a simple database and data-processing architecture used in the early days of computing and is characterized by high-volume and -dimensional data that is rapidly updated represents a phenomenon or feature that has emerged in the digital age. Across the medical industry, various types of medical data are generated at a high speed, and trends indicate that applying big data in the medical field helps improve the quality of medical care and optimizes medical processes and management strategies [2, 3]. Currently, this trend is shifting from civilian medicine to military medicine. For example, the United States is exploring the potential to use of one of its largest healthcare systems (the Military Healthcare System) to provide healthcare to eligible veterans in order to potentially benefit > 9 million eligible personnel [4]. Another data-management system has been developed to assess the physical and mental health of active-duty personnel, with this expected to yield significant economic benefits to the military medical system [5]. However, in medical research, the wide variety of clinical data and differences between several medical concepts in different classification standards results in a high degree of dimensionality heterogeneity, timeliness, scarcity, and irregularity to existing clinical data [6, 7]. Furthermore, new data analysis techniques have yet to be popularized in medical research [8]. These reasons hinder the full realization of the value of existing data, and the intensive exploration of the value of clinical data remains a challenging problem.
Computer scientists have made outstanding contributions to the application of big data and introduced the concept of data mining to solve difficulties associated with such applications. Data mining (also known as knowledge discovery in databases) refers to the process of extracting potentially useful information and knowledge hidden in a large amount of incomplete, noisy, fuzzy, and random practical application data [9]. Unlike traditional research methods, several data-mining technologies mine information to discover knowledge based on the premise of unclear assumptions (i.e., they are directly applied without prior research design). The obtained information should have previously unknown, valid, and practical characteristics [9]. Data-mining technology does not aim to replace traditional statistical analysis techniques, but it does seek to extend and expand statistical analysis methodologies. From a practical point of view, machine learning (ML) is the main analytical method in data mining, as it represents a method of training models by using data and then using those models for predicting outcomes. Given the rapid progress of data-mining technology and its excellent performance in other industries and fields, it has introduced new opportunities and prospects to clinical big-data research [10]. Large amounts of high quality medical data are available to researchers in the form of public databases, which enable more researchers to participate in the process of medical data mining in the hope that the generated results can further guide clinical practice.
This article provided a valuable overview to medical researchers interested in studying the application of data mining on clinical big data. To allow a clearer understanding of the application of data-mining technology on clinical big data, the second part of this paper introduced the concept of public databases and summarized those commonly used in medical research. In the third part of the paper, we offered an overview of data mining, including introducing an appropriate model, tasks, and processes, and summarized the specific methods of data mining. In the fourth and fifth parts of this paper, we introduced data-mining algorithms commonly used in clinical practice along with specific cases in order to help clinical researchers clearly and intuitively understand the application of data-mining technology on clinical big data. Finally, we discussed the advantages and disadvantages of data mining in clinical analysis and offered insight into possible future applications.
A public database describes a data repository used for research and dedicated to housing data related to scientific research on an open platform. Such databases collect and store heterogeneous and multi-dimensional health, medical, scientific research in a structured form and characteristics of mass/multi-ownership, complexity, and security. These databases cover a wide range of data, including those related to cancer research, disease burden, nutrition and health, and genetics and the environment. Table 1 summarizes the main public medical databases [11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26]. Researchers can apply for access to data based on the scope of the database and the application procedures required to perform relevant medical research.

A previous study identified sepsis as a major cause of death in ICU patients [100]. The authors noted that the predictive model developed previously used a limited number of variables, and that model performance required improvement. The data-mining process applied to address these issues was, as follows: (1) data selection using the MIMIC III database; (2) extraction and integration of three types of data, including multivariate features (demographic information and clinical biochemical indicators), time series data (temperature, blood pressure, and heart rate), and clinical latent features (various scores related to disease); (3) data cleaning and transformation, including fixing irregular time series measurements, estimating missing values, deleting outliers, and addressing data imbalance; (4) data mining through the use of logical regression, generation of a decision tree, application of the RF algorithm, an SVM, and an ensemble algorithm (a combination of multiple classifiers) to established the prediction model; (5) pattern evaluation using sensitivity, precision, and the area under the receiver operating characteristic curve to evaluate model performance; and (6) evaluation of the results, in this case the potential to predicting the prognosis of patients with sepsis and whether the model outperformed current scoring systems.
Wu et al. [101] noted that traditional survival-analysis methods often ignored the influence of competitive risk events, such as suicide and car accident, on outcomes, leading to deviations and misjudgements in estimating the effect of risk factors. They used the SEER database, which offers cause-of-death data for cancer patients, and a competitive risk model to address this problem according to the following process: (1) data were obtained from the SEER database; (2) demography, clinical characteristics, treatment modality, and cause of death of cecum cancer patients were extracted from the database; (3) patient data were deleted when there were no demographic, clinical, therapeutic, or cause-of-death variables; (4) Cox regression and two kinds of competitive risk models were applied for survival analysis; (5) the results were compared between three different models; and (6) the results revealed that for survival data with multiple endpoints, the competitive risk model was more favourable.
A study by Martínez Steele et al. [102] applied PCA for nutritional epidemiological analysis to determine dietary patterns and evaluate the overall nutritional quality of the population based on those patterns. Their process involved the following: (1) data were extracted from the NHANES database covering the years 2009–2010; (2) demographic characteristics and two 24 h dietary recall interviews were obtained; (3) data were weighted and excluded based on subjects not meeting specific criteria; (4) PCA was used to determine dietary patterns in the United States population, and Gaussian regression and restricted cubic splines were used to assess associations between ultra-processed foods and nutritional balance; (5) eigenvalues, scree plots, and the interpretability of the principal components were reviewed to screen and evaluate the results; and (6) the results revealed a negative association between ultra-processed food intake and overall dietary quality. Their findings indicated that a nutritionally balanced eating pattern was characterized by a diet high in fibre, potassium, magnesium, and vitamin C intake along with low sugar and saturated fat consumption.
The use of “big data” has changed multiple aspects of modern life, with its use combined with data-mining methods capable of improving the status quo [86]. The aim of this study was to aid clinical researchers in understanding the application of data-mining technology on clinical big data and public medical databases to further their research goals in order to benefit clinicians and patients. The examples provided offer insight into the data-mining process applied for the purposes of clinical research. Notably, researchers have raised concerns that big data and data-mining methods were not a perfect fit for adequately replicating actual clinical conditions, with the results potentially capable of misleading doctors and patients [86]. Therefore, given the rate at which new technologies and trends progress, it is necessary to maintain a positive attitude concerning their potential impact while remaining cautious in examining the results provided by their application.
In the future, the healthcare system will need to utilize increasingly larger volumes of big data with higher dimensionality. The tasks and objectives of data analysis will also have higher demands, including higher degrees of visualization, results with increased accuracy, and stronger real-time performance. As a result, the methods used to mine and process big data will continue to improve. Furthermore, to increase the formality and standardization of data-mining methods, it is possible that a new programming language specifically for this purpose will need to be developed, as well as novel methods capable of addressing unstructured data, such as graphics, audio, and text represented by handwriting. In terms of application, the development of data-management and disease-screening systems for large-scale populations, such as the military, will help determine the best interventions and formulation of auxiliary standards capable of benefitting both cost-efficiency and personnel. Data-mining technology can also be applied to hospital management in order to improve patient satisfaction, detect medical-insurance fraud and abuse, and reduce costs and losses while improving management efficiency. Currently, this technology is being applied for predicting patient disease, with further improvements resulting in the increased accuracy and speed of these predictions. Moreover, it is worth noting that technological development will concomitantly require higher quality data, which will be a prerequisite for accurate application of the technology.
Finally, the ultimate goal of this study was to explain the methods associated with data mining and commonly used to process clinical big data. This review will potentially promote further study and aid doctors and patients.
Biologic Specimen and Data Repositories Information Coordinating Center
China Health and Retirement Longitudinal Study
China Health and Nutrition Survey